Note that the fragment ions containing lysines 5, 8, or 12 show a similar ratio of the acetylated (Ac) form to the nonacetylated (propionylated ) form, whereas the peptide containing K16 (middle panel) is strongly hypoacetylated in cells treated with siRNA against hMOF. (E) Enlarged regions of the MS/MS spectra generated from the monoacetylated peptide from control (left panels) or hMOF knockdown (right panels) cells. To analyze a specific effect of hMOF knockdown on H4K16 acetylation, we have analyzed the monoacetylated peptide by MS/MS. (D) Schematic display of the fragment ions generated by collision-induced decay from a monoacetylated peptide. For quantitation, H4 molecules have been modified using D6 acetic anhydride rather than propionic anhydride to ensure similar ionization behavior of the in vitro- and in vivo-modified peptides. (C) Quantitation of acetylation levels seen in vivo. Note that the peptide with the highest m / z carries four propionyl groups and corresponds to the unmodified form in vivo. Only the peptide containing amino acids 4 to 17 of H4 is shown.
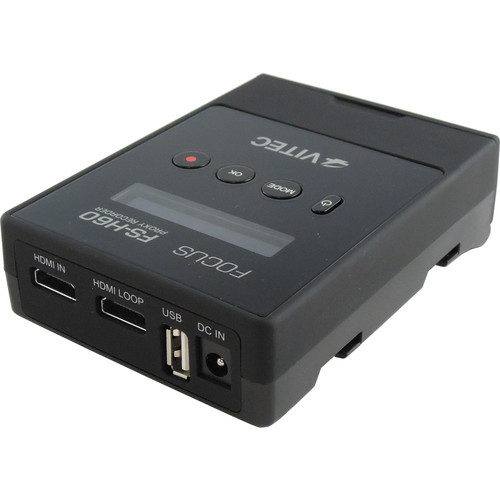

(B) MALDI-time of flight analysis of H4 molecules purified via SDS-PAGE and propionylated and digested as described previously (6). (Left panel) Western blot analysis to show reduction of proteins levels by MOF siRNA 1 and 2 in comparison to control siRNA. The amount of histones loaded was controlled by staining the membrane with Ponceau S. Western blot analysis of the acid-soluble fraction was performed with acetylation-specific antibodies as indicated. Cells were transfected with either control or hMOF-1 siRNA. (A, right and middle panels) Acetylation of histone H4 lysine 16 is specifically reduced in hMOF-depleted cells. Right panel: detection of modified residues with acetylation-specific antibodies.Īnalysis of the histone modification state in cells treated with siRNA against hMOF. Left panel: autoradiograph (top) and corresponding Coomassie gel (bottom). (E) Nucleosomal HAT activity of anti-FLAG immunoprecipitates from HA–2 ϫ FLAG-tagged hMOF or from HeLa cells. Right panel: comparison of hMOF (lane 10) and dMOF (lane 11) HAT activity on nucleosomes. Bottom, Coomassie staining of the same gel. Left panel: top and middle, autoradiographs (short exposure, overnight long exposure, 2 weeks) of the gel. (D) Separation of histones from HAT assay by SDS-PAGE. Bars 1 to 4 correspond to lanes 1 to 4 in panel B bar 5, GST alone bar 6, histones alone. (C) Filter binding HAT assay with different hMOF constructs using a recombinant histone octamer as a substrate for acetylation. (B) Constructs of GST-hMOF and FLAG-hMS元 expressed in E. (A) Schematic representation of the GST-hMOF fusion constructs used in histone acetyltransferase assays. Taken together, our data show that hMOF is requiredįor histone H4 lysine 16 acetylation in mammalian cells and suggest that hMOF has a role in DNA damage response during cell

Furthermore, hMOF-depleted cells show an increased number of phospho-ATMĪnd γH2AX foci and have an impaired repair response to ionizing radiation.
Treatment with specific inhibitors of the DNA damage response pathway reverts the cell cycleĪrrest caused by a reduction in hMOF protein levels. Protein levels by RNA interference in HeLa cells also leads to accumulation of cells in the G2 and M phases of the cell cycle. hMOF and hMS元 small interfering RNA-treatedĬells also show dramatic nuclear morphological deformations, depicted by a polylobulated nuclear phenotype. We also provide evidence that, similar to the Drosophila dosage compensation system, hMOF and hMS元 form a complex in mammalian cells. Knockdown of hMOF in HeLa and HepG2 cells causes a dramatic reduction of histone H4K16 acetylation as detected by Westernīlot analysis and mass spectrometric analysis of endogenous histones. hMOF is also required for this modification in mammalianĬells. That the human orthologue of Drosophila melanogaster MOF, hMOF, is a histone H4 lysine K16-specific acetyltransferase. Reversible histone acetylation plays an important role in regulation of chromatin structure and function.
